Quick Links
- QB Graduate Program Faculty Members
- Student Members
QB Graduate Program Faculty and Students
IQB Members come from virtually all of Rutgers STEM-related schools and centers, including the Schools of Arts and Sciences (SAS), Engineering (SOE), Environmental and Biological Sciences (SEBS), Health Professions (SHP), the Ernest Mario School of Pharmacy (EMSOP), the Graduate School-Newark (GS-N), the New Jersey Medical School (NJMS), the Robert Wood Johnson Medical School (RWJMS), the Cancer Institute of New Jersey (CINJ), the Center for Advanced Biotechnology and Medicine (CABM), the Child Health Institute of New Jersey (CHINJ), the Environmental and Occupational and Health Sciences Institute (EOHSI), the Human Genetics Institute of New Jersey (HGINJ/RUCDR Infinite Biologics), the New Jersey Institute for Food, Nutrition, and Health (NJIFNH), the Waksman Institute of Microbiology (WIM), and the Office of Research and Economic Development (ORED).

| Gabriela Alexe Adjunct Faculty Broad Institute One research direction is Integrative Cancer Genomics, where I am working on extracting and creating knowledge from multiple genomic data platforms to explore the global mechanisms of cancer initiation, progression and metastasis, with a particular emphasis on discovering driver genetic events and combinatorial biomarkers for improving risk profiling, therapy specificity and treatment efficacy.
The second research direction is Cancer Immunology, where I am working on deciphering the mechanisms associated with T cell dysfunction and immune signatures in cancer.
|
Dana-Farber Cancer Institute
Room 610 616-632-6782
|

| David Alland Professor Medicine, Rutgers New Jersey Medical School Chief, Infectious Diseases Resident Member, Public Health Research Institute Associate Dean, Clinical Research The Alland laboratory is interested in many different aspects of M. tuberculosis molecular biology, epidemiology and diagnostics. Many of these areas are related to understanding how antibiotic resistance develops on both the cellular and epidemiological level. We also have a major program in biodefense diagnostics and pathogenesis.
|
Medical Sciences Building
Room A920C 973-972-9834
|

| Ioannis Androulakis Professor and Undergraduate Program Director Biomedical Engineering, School of Engineering Our research interests focus on the quantitative systems biology and quantitative systems pharmacology of inflammation and the interplay between immune response and biological rhythms. We are interested in delineating the intricacies of the human response to stress as well as characterizing responses to drugs, including the opportunities for personalizing oral drug formulations to target groups according to sex, age and disease status.
|
Biomedical Engineering
Room 212 848-445-6561
|

| Eddy Arnold Board of Governors Professor Chemistry & Chemical Biology, School of Arts & Sciences Resident Member, Center for Advanced Biotechnology & Medicine Full Member, Rutgers Cancer Institute of New Jersey Drug design and development targeting HIV/AIDS and flu, and structural biology of HIV-1 reverse transcriptase and integrase, flu polymerase, and bacterial RNA polymerases. A key focal area is to define the structural basis of drug resistance in infectious diseases, and to use structural information to design drugs to overcome drug resistance, including the development of generally applicable strategies.
|
Center for Advanced Biotechnology & Medicine
Room 016 848-445-9777
|

| Gary Aston-Jones Director, Brain Health Institute Murray and Charlotte Strongwater Endowed Chair in Neuroscience & Brain Health Rutgers Biomedical & Health Sciences Brain neuromodulatory systems, cognitive performance, drug abuse, sleep and waking, and affective disorders
|
Rutgers Brain Health Institute
Room 259A 732-235-6077
|

| David E. Axelrod Professor Genetics, School of Arts & Sciences Our lab is interested in developing quantitative prognostic factors for detecting early stages of breast and colon cancer, stratifying patients for good and poor survival, and suggesting improved treatments of patients with cancer. We combine experimental techniques such as image analysis of human biopsy specimens to measure the expression of oncogenes and tumor suppressor genes and proteins; and mathematical and computer modeling to develop agent based models of normal stem cell stochastic dynamics, abnormal proliferation and development, initiation of cancer, progression to invasive cancer, tumor heterogeneity, and chemoprevention.
|
Nelson Biology Laboratories
Room B341 848-445-2011
|

| William Belden Associate Professor Animal Sciences, School of Environmental & Biological Sciences The research in my laboratory focuses on understanding the molecular aspects of chromatin-remodeling and circadian rhythms. My lab uses the highly-tractable model eukaryote Neurospora crassa in combination with biochemical, genetic and genome-wide studies to understand the molecular mechanisms of chromatin-remodeling in clock function.
|
Foran Hall
Room 326 848-932-5617
|

| Helen M. Berman Board of Governors Distinguished Professor Emerita Chemistry & Chemical Biology, School of Arts & Sciences Director Emerita, RCSB Protein Data Bank Founding Director, Center for Integrative Proteomics Research Resident Member, Institute for Quantitative Biomedicine From 1998-2014, I was the Director of the Research Collaboratory for Structural Bioinformatics Protein Data Bank (RCSB PDB). RCSB PDB is a member of the Worldwide Protein Data Bank (wwPDB) that manages the PDB archive of information about the structures of proteins, nucleic acids, and complex assemblies. I play a leadership role for the EMDataBank and am working to develop infrastructure for the archiving of structures determined using Integrative/Hybrid Methods.
The work on these projects has been informed by our research and studies in structural biology, in particular protein-nucleic acid interactions, collagen, binary and ternary complexes with catabolite activating protein (CAP), and nucleic acids. Currently, we are working in new classification schemes for transcription factors with a focus on understanding the details of the protein-nucleic acid interface. Most recently I am working on how to use film to communicate to a broader audience about the importance of structural biology in medicine and health. |
Proteomics Building
Room 208NP 848-445-4667
|

| Debashish Bhattacharya Distinguished Professor Ecology, Evolution & Natural Resources, School of Environmental & Biological Sciences Work in the Bhattacharya lab primarily addresses the origin and evolution of photosynthetic eukaryotes. We use a variety of techniques ranging from algal culture and physiology, standard and single cell genomics, and functional genomics to study how algae gained and orchestrate the functions of their photosynthetic organelle, the plastid and how they adapt to changing environmental conditions. We have recently initiated a large-scale genomic and functional genomic analysis of corals and their symbionts to understand the basis of biomineralization, the genetic structure of coral populations, and their pathways of stress tolerance.
|
Foran Hall
Room 102 848-932-6218
|

| Kenneth J. Breslauer Linus C. Pauling Professor Chemistry & Chemical Biology, School of Arts & Sciences Dean, Biological Sciences Vice President, Health Science Partnerships The research conducted in my laboratory combines both biophysical and bioorganic chemistry to investigate the following four interrelated programs of research.
|
Wright Rieman Laboratories
Room 155 732-445-3954
|
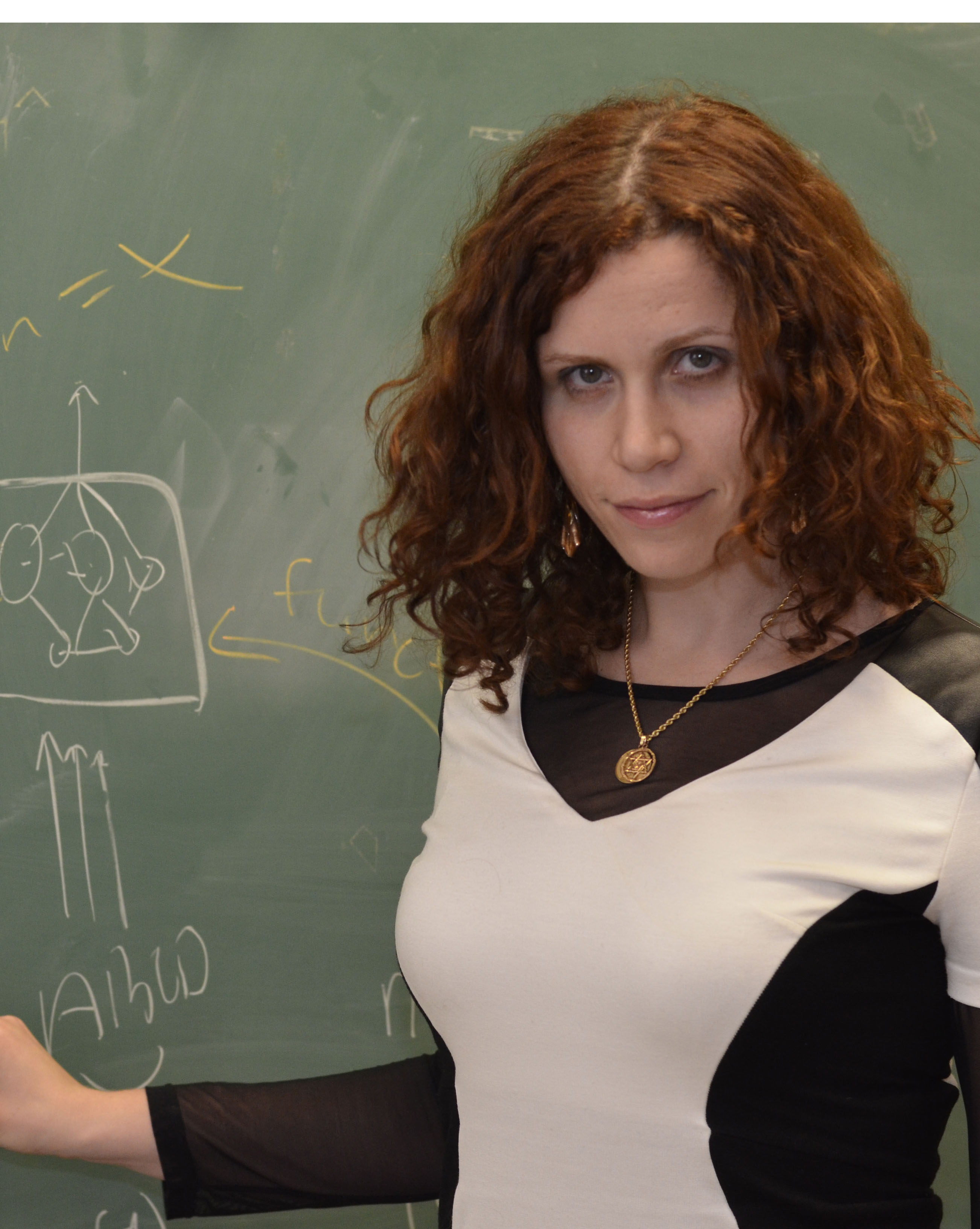
| Yana Bromberg Associate Professor Biochemistry & Microbiology, School of Environmental & Biological Sciences Bioinformatics approaches to protein function prediction and genome variation analysis. Our main goal is to develop fast, accurate, and meaningful ways of analyzing this growing deluge of biological data and to bring these developments bench- (or patient-) side. To make our predictions we rely on a number of sequence-based features (including evolutionary information and other predictor results) and utilize a variety of methodologies (including Neural Nets, SVMs and random forests).
|
Lipman Hall
Room 218 848-932-5638
|

| Linda Brzustowicz Distinguished Professor & Chair Genetics, School of Arts & Sciences Full Academic Member, Human Genetics Institute of New Jersey My research group applies the techniques of molecular and statistical genetics to approach clinically relevant problems in neuroscience, with the ultimate goal of understanding gene function in both the pathologic and normal states. We are currently studying schizophrenia, autism, and specific language impairment (SLI). Work directly conducted by my group includes development of phenotype definitions, subject recruitment and assessment (for autism and SLI), genotyping and statistical analysis for linkage and association studies, comparative genomic analysis, and gene expression studies.
|
Life Sciences
Room 231 848-445-1638
|

| Stephen K. Burley, M.D., D.Phil. University Professor & Henry Rutgers Chair Founding Director, Institute for Quantitative Biomedicine Director, RCSB Protein Data Bank Chemistry & Chemical Biology, School of Arts & Sciences Member and Cancer Pharmacology Program Co-Lead, Rutgers Cancer Institute of New Jersey As University Professor and Henry Rutgers Chair, Founding Director of the
Institute for Quantitative Biomedicine at Rutgers, The State University of New Jersey, and a Member of The Cancer Institute of New Jersey, I aim to foster interactive networks of research groups that utilize multidisciplinary approaches to address important biological and biomedical challenges.
In my capacity as Director of the RCSB Protein Data Bank, dividing my time between Rutgers and the University of California at San Diego, I work to sustain open access to the single global archive of biomolecular structure data with >99% uptime 24/7/365. |
Proteomics Building Room 117
|

| Javier Cabrera Professor Statistics & Biostatistics, School of Arts & Sciences Rutgers Cardiovascular Institute New methodology for the analysis of big data in medicine, genomics, clinical and registry data.
|
Hill Center Room 471
848-445-7665
|

| David Case Distinguished Professor Chemistry & Chemical Biology, School of Arts & Sciences Resident Member, Institute for Quantitative Biomedicine Our group studies theoretical chemistry of biomolecules, using molecular dynamics simulations of proteins and nucleic acids, electronic structure calculations of transition-metal complexes that model active sites in metalloenzymes, and development and application of methods for NMR structure determination.
|
Proteomics Building Room 208AB
848-445-5885
|

| Chang S. Chan Assistant Professor Medicine, Robert Wood Johnson Medical School Resident Member, Rutgers Cancer Institute of New Jersey Associate Director, Center for Systems & Computational Biology High-throughput sequencing and integration of data in the study of genetics of cancer, complex diseases, and gene regulation, and Integration of diverse data with phenotypes for a systems level understanding of biology
|
Rutgers Cancer Institute of New Jersey
732-235-7363
|

| Sunita Chaudhary Adjunct Assistant Professor Surgery, Robert Wood Johnson Medical School Director, Research Education Resident Member, Rutgers Cancer Institute of New Jersey Our current research focuses on understanding the influence of race, gender and immigration status on biomedical career choice.
|
Rutgers Cancer Institute of New Jersey
Room 8005 732-235-9869
|

| Neeraj Chauhan Assistant Professor Microbiology & Molecular Genetics, Rutgers New Jersey Medical School Resident Member, Public Health Research Institute The focus of my current research is the opportunistic human pathogenic yeast, Candida albicans. It is the leading cause of invasive fungal disease in premature infants, surgical patients and cancer patients receiving immunosuppressive chemotherapy. My research is directed to elucidate the complex signaling pathways that contribute to the virulence & pathogenesis of this organism. Our approaches include molecular, biochemical and immunological techniques to study these events. This involves isolation of encoding genes, construction of knock-out strains to study gene function, DNA Microarrays, and GFP localization etc. Two component signaling proteins (histidine kinases and response regulators) and downstream MAP kinases are the main focus of research.
|
Public Health Research Institute
973-854-3470
|

| Shishir Chundawat Associate Professor Chemical & Biochemical Engineering, School of Engineering The Chundawat Research Group takes a carbohydrate or ‘glycan-centric’ approach to develop advanced protein and glycan engineering (or broadly glycoengineering) toolkits along with applying novel bioprocessing and biophysical techniques to address fundamental scientific and engineering problems relevant to healthcare, bioenergy, and biomaterials research. He has multidisciplinary expertise working with carbohydrate-active enzymes (CAZymes); protein modeling and engineering; carbohydrate chemistry; biomanufacturing; and developing novel analytical techniques for characterization of glycans and protein/CAZymes-glycan interactions. His team at Rutgers is actively working towards expanding the repertoire of applications for carbohydrate-active enzymes and glycans in general in the area of biomanufacturing and human health.
|
School of Engineering
Room C-150A 848-445-3678
|

| William Craelius Professor Biomedical Engineering, School of Engineering In my lab, we study how technology can restore mobility to persons with missing or impaired limbs. My research with amputees developed the first prosthesis that restored multiple finger dexterity. Since then I have translated this technology to assist those with paralysis due to brain injuries. My work with many clients having arm paralysis due to stroke, brain injury and cerebral palsy, has taught me both the potential and limitations of tools that attempt to promote neuroplastic recovery in these populations. I employ technology to measure arm motion and hand control, using cameras and various sensors, and have demonstrated their therapeutic potential. We are also developing new growth substrates for helping nerves regenerate.
|
Biomedical Engineering
Room 207 848-445-6558
|

| Wei Dai Assistant Professor Cell Biology & Neuroscience, School of Arts & Sciences Resident Member, Institute for Quantitative Biomedicine Associate Member, Rutgers Cancer Institute of New Jersey Our research focuses on charactering the structures of macromolecular machinery in their cellular context using three-dimensional (3D) cryo-electron tomography (cryoET) and correlative light and electron microscopy. Structures of intracellular macromolecular complexes are usually heterogeneous and dynamic, relying largely on interactions with other cellular components. Using cryoET to determine the structure of cellular machinery inside the cell avoids damage to the complexes during purification, captures snapshots of the complex while in action, and provides information on cross-talk of the complexes with their cellular partners during biological processes. Our current research interest includes: Huntington's disease: structure and organization of protein aggregates; 3D architecture of the diseased cells under misfolded protein aggregation stress & Structural biology of phage/virus maturation.
|
Proteomics Building Room 208G
848-445-6560
|

| Tirthankar Dasgupta Associate Professor Causal inference, experimental design: foundations, philosophies, methodologies, and applications to physical, social, biomedical and engineering sciences, sequential exploration of complex surfaces, statistical modeling of complex physical and engineering systems, quality engineering and statistical process control.
|
Hill Center Room 501
848-445-7271
|

| Subhajyoti De Assistant Professor Pathology & Laboratory Medicine, Robert Wood Johnson Medical School Systems & Computational Biology Resident Member, Rutgers Cancer Institute of New Jersey The De laboratory develops and applies genomics, systems biology, and mathematical modeling approaches to understand the biology of cancer and use that knowledge for better diagnosis, stratification of cancer patients and their personalized treatment.
|
Rutgers Cancer Institute of New Jersey
732-235-8558
|

| Monica Driscoll Distinguished Professor Molecular Biology & Biochemistry, School of Arts & Sciences Our lab uses the elegant and powerful model system C. elegans to decipher conserved molecular mechanisms of cellular function and dysfunction. The main problems we investigate are neuronal degeneration, neuronal regeneration, and the biology of aging, with a focused goal of defining strategies for extending healthspan.
|
Nelson Biology Laboratories
Room A232 848-445-7182
|

| Siobain Duffy Associate Professor Ecology, Evolution & Natural Resources, School of Environmental & Biological Sciences The Duffy lab uses a blend of computational and experimental approaches to understand and model the evolution of emerging RNA and single-stranded DNA viruses.
|
Foran Hall
Room 316 848-932-6299
|

| Shuchismita Dutta Scientific Educational Development Lead, RCSB Protein Data Bank Associate Research Professor, Institute for Quantitative Biomedicine I am a structural biologist, dedicated to promoting a molecular structural view of biology and medicine. I work with educators to develop curricular materials and case studies for students (high school, undergraduate, and graduate) to learn about biomolecular structures and visualization. I am also committed to STEM Education, and interested in Science Pedagogy.
|
Institute for Quantitative Biomedicine
Room 114 848-445-4915
|

| Richard H. Ebright Board of Governors Professor Chemistry & Chemical Biology, School of Arts & Sciences Principal Investigator, Waksman Institute of Microbiology Richard H. Ebright seeks to understand structures, mechanisms, and regulation of bacterial transcription complexes and to identify, characterize, and develop small-molecule inhibitors of bacterial transcription for application as antituberculosis agents and broad-spectrum antibacterial agents.
|
Waksman Institute of Microbiology
Room 201-A 848-445-5179
|

| Christopher Ellison Assistant Professor Genetics, School of Arts & Sciences 3D genome organization and transposable element evolution in Drosophila
|
Nelson Biological Laboratories
Room B420 848-445-2812
|

| Laura Fabris Associate Professor Materials Science & Engineering, School of Engineering We are interested in the synthesis and characterization of plasmonic nanoparticles and their use in surface enhanced Raman scattering (SERS)-based sensing and cell imaging, and in those applications that require near field enhancement effects. We also focus on the study of nanoparticle assembly in solution and on unraveling the fundamental phenomena at the basis of ligand exchange in plasmonic nanoparticles in solution.
|
McLaren Center for Ceramic Research
Room 216 848-445-5606
|

| Mikolai Fajer Senior Scientist Schrodinger, Inc. Free energy and conformer sampling of protein computational models
|
|

| Paul Falkowski Board of Governors Professor & Bennett L. Smith Chair, Business & Natural Resources Earth & Planetary Sciences and Marine & Coastal Sciences, School of Environmental & Biological Sciences Director, Rutgers Energy Institute My research interests are focused on three areas - origins of life, how electron transfer reactions are mediated, and how organisms transformed the geochemistry of Earth. In the evolution of Earth, microbes became a major force in transforming this planet to make it habitable for animals, including humans. I seek to understand the basic chemical reactions that enabled microbes to transform Earth's goechemistry. I work at the molecular level of proteins and fundamental chemical reactions of minerals, and the global scale of how this planet came to have oxygen as the second most abundant gas. I am most interested in understanding how these kinds of processes have transformed our planet and may evolve on planetary bodies in our solar system and on extra-solar planets. There are only two questions I address: Where did we come from? And are we alone?
|
Institute of Marine & Coastal Sciences
Room 318D 848-932-3426
|

| Zukang Feng Research Professor at IQB RCSB Protein Data Bank My research interest focuses on developing tools for data acquisition, data validation, data standardization, and data analysis in the structural biology.
|
Proteomics Building
Room 113 |

| David J. Foran Professor Pathology and Laboratory Medicine; Radiology, Robert Wood Johnson Medical School Director, Center for Biomedical Imaging Resident Member, Rutgers Cancer Center of New Jersey A major concentration for my laboratory has been the development of a family of data-mining, imaging and computational tools for characterizing a wide range of malignancies and elucidating the role that protein and molecular expression plays in disease onset and progression. My training, experience and expertise gained while leading and supporting multi-investigator projects in computational imaging, computer-assisted diagnostics, bioinformatics, team-based software development and high-performance computing have provided me with the requisite tools and skills to work in partnership with basic, clinical and translational researchers to address fundamental problems in cancer detection, patient stratification, disease management, and outcomes studies.
|
Rutgers Cancer Institute of New Jersey
Room 3552 732-235-6925
|

| Joel Freundlich Associate Professor Pharmacology, Physiology & Neuroscience, Rutgers New Jersey Medical School The Freundlich group is a chemical biology lab that utilizes a multi-disciplinary approach to study infectious diseases, with a specific focus on tuberculosis. Within the Department of Pharmacology & Physiology and the Department of Medicine (Division of Infectious Diseases, Center for Emerging & Re-emerging Pathogens), we focus on 1) the development and application of novel computational methods to discover and optimize chemical probes with which to study the pathogenesis of disease and 2) the utilization of biological techniques to elucidate how each probe affects disease pathogenesis, often through the complex modulation of more than one biological target. We assert that this approach has significant potential to seed the discovery of novel biological targets and small molecule drug discovery hits/leads.
|
Medical Sciences Building
Room I-503 973-972-7165
|

| Zoran Gajic Professor Electrical & Computer Engineering, School of Engineering Graduate Program Director Research interests are in controls systems, energy systems (fuel and solar cells, wind, smart grids), wireless communications, and networking.
|
Electrical Engineering
Room 134A 732-445-3415
|

| Shridar Ganesan Associate Professor & Omar Boraie Chair, Genomic Science Medicine & Pharmacology, Robert Wood Johnson Medical School Resident Member, Rutgers Cancer Institute of New Jersey Section Chief, Molecular Oncology Associate Director, Translational Science With a research interest in breast cancer biology and DNA repair, I am currently exploring how DNA repair defects in cancers can be exploited to develop novel effective treatments. I am also active in applying next-generation sequencing technology to identify specific genomic changes in cancers that can be therapeutically targeted. As a physician/scientist I both run a basic research laboratory focused on breast cancer biology and see patients in the Stacy Goldstein Breast Cancer Center. In the clinic, I work collaboratively with experts across multi-disciplines and have the opportunity to put theory into practice as we aim to develop the next generation of targeted treatments for breast cancer. Working with a team of radiation oncologists, surgical oncologists, nurses, social workers, genetic specialists and others, I help patients understand their specific disease and their treatment options so that they can make informed decisions. I am also an associate professor of medicine and pharmacology at Robert Wood Johnson Medical School.
|
Rutgers Cancer Institute of New Jersey
732-235-7428
|

| Marc R. Gartenberg Professor Biochemistry & Molecular Biology, Robert Wood Johnson Medical School Full Member, Rutgers Cancer Institute of New Jersey My lab investigates the ways that heterochromatin is assembled, maintained and inherited. Our recent work includes focus on the specialized roles of heterochromatin in sister chromatid cohesion. We use budding yeast as a model eukaryotic system where we have developed molecular biological tools to ask questions at the mechanistic level.
|
Robert Wood Johnson Medical School Research Building/School of Public Health
Room 284 732-235-5800
|

| Michael Gatza Assistant Professor Radiation Oncology, Robert Wood Johnson Medical School Resident Member, Rutgers Cancer Institute of New Jersey Our research program is focused on understanding the mechanisms of oncogenic signaling and therapeutic response in human breast and ovarian cancers. Leveraging a multi-disciplinary approach spanning cancer biology, molecular biology, genomics, genetics, bioinformatics, and statistics allows us to identify relevant genomic alterations in human tumors and then experimentally investigate the mechanisms by which these alterations affect given tumor phenotypes in order to develop rational therapeutic strategies based on the unique complement of genomic alterations present in each patient's tumor.
|
Rutgers Cancer Institute of New Jersey
Room 4558 732-235-8751
|

| Panos Georgopoulos Professor Rutgers Biomedical & Health Sciences Director, Ozone Research Center Multiscale computational modeling of environmental and biological systems and of their interactions; computational toxicology; quantitative frameworks for adverse outcome pathways in environmental health systems.
Enviroinformatics, bioinformatics, and socioinformatics for public health applications; predictive data analytics for human exposures and associated health outcomes. |
Environmental & Occupational Health Sciences Institute
Room 308 848-445-0159
|

| Nasrin Ghesani Associate Professor Radiology, Rutgers New Jersey Medical School Molecular imaging and Positron Emission Tomography
|
University Hospital, Newark
Room UH 141 973-972-1770
|
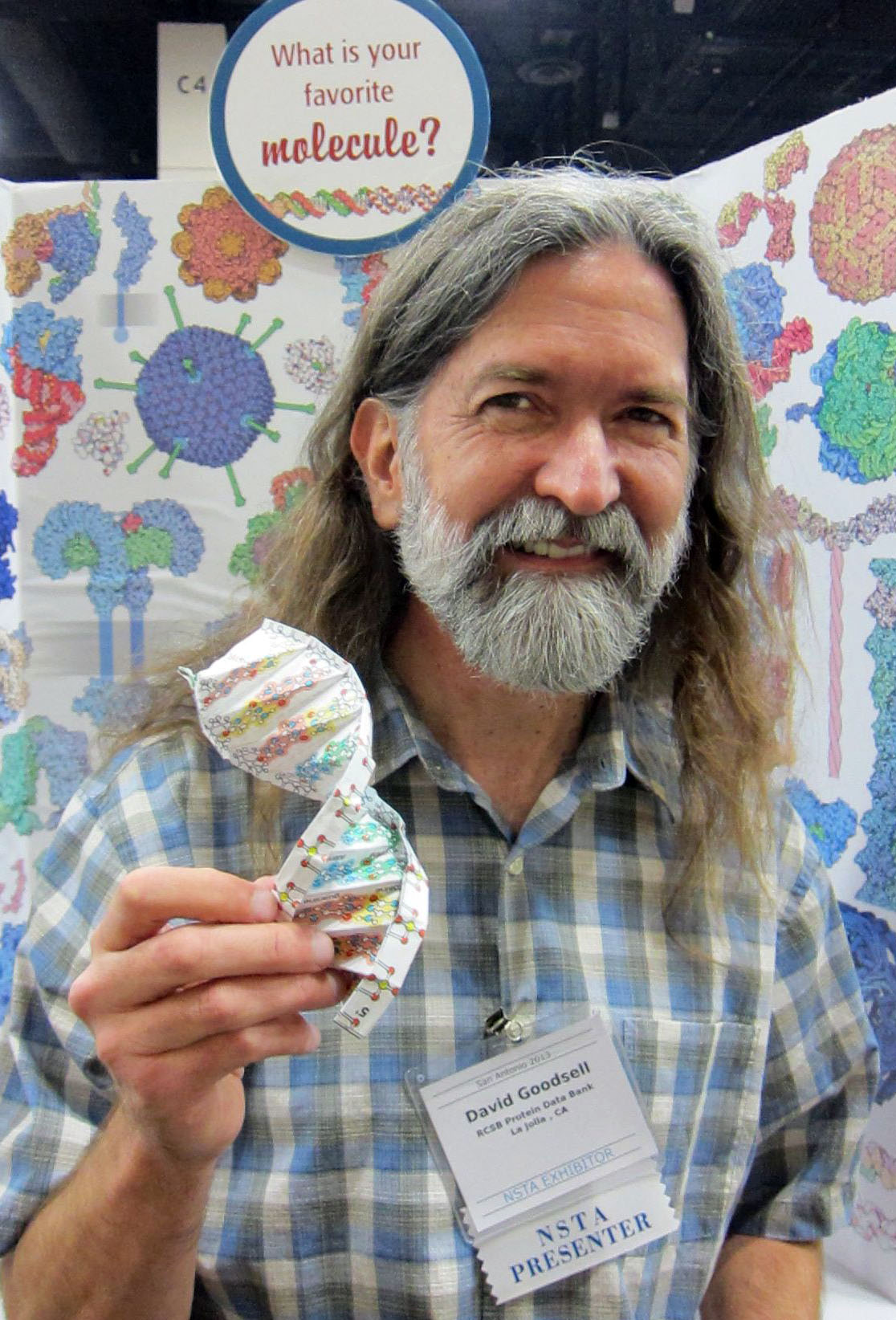

| David Goodsell Associate Professor Molecular Biology, The Scripps Research Institute Research Professor, Chemistry & Chemical Biology, Rutgers University My research uses computer graphics and simulation to explore structure/function relationships in key biological systems. For two decades, I have pioneered approaches for the integrative modeling of the cellular mesoscale, visualizing the molecular structure of cellular environments and entire cells. Building on AutoDock, the first method for computational docking of flexible molecules to proteins and DNA, I continue to develop advanced docking methods and apply them to the understanding of HIV and its interaction with cells of the immune system. Science education is also a strong focus of my laboratory. I am author of the Molecule of the Month, a feature at the RCSB Protein Data Bank that presents the structure and function of a new molecule each month, and several illustrated books on biological molecules, their diverse roles within living cells, and the growing connections between biology and nanotechnology.
|
The Scripps Research Institute
858-784-2839
|

| Derek Gordon Associate Professor Department of Genetics, School of Arts & Sciences My research focuses on linkage and associate methods. I perform both theoretical and applied studies of these methods. My theoretical work is mainly concerned with phenotype and genotype misclassification and their effects on tests of linkage and association. My applied work involves gene mapping studies of scoliosis, Alzheimer’s Disease, and hair and skin phenotypes
|
Nelson Biological Laboratories Room C213
848-445-3386
|

| Masayori Inouye Distinguished Professor Biochemistry & Molecular Biology, Robert Wood Johnson Medical School Molecular biology of cellular adaptation to stresses
|
Center for Advanced Biotechnology & Medicine
Room 140 848-445-9813
|

| Kenneth Irvine Distinguished Professor Molecular Biology & Biochemistry, School of Arts & Sciences Principal Investigator, Waksman Institute of Microbiology The lab is engaged in projects whose long-term goal is to define relationships between patterning and growth in developing and regenerating organs. Much of their research takes advantage of the powerful genetic, molecular, and cellular techniques available in Drosophila melanogaster, which facilitate both gene discovery and the analysis of gene function.
|
Waksman Institute of Microbiology
Room 0136 848-445-3382
|

| Mehdi Javanmard Assistant Professor Electrical & Computer Engineering, School of Engineering Our lab focuses on exploiting emerging micro- and nanotechnologies for health and environmental monitoring.
|
Electrical & Computer Engineering Room 211
848-445-3382
|

| Shantenu Jha Associate Professor Electrical & Computer Engineering, School of Engineering High-performance and distributed computing; computational and data-intensive science and engineering
|
Electrical Engineering
Room 211 |
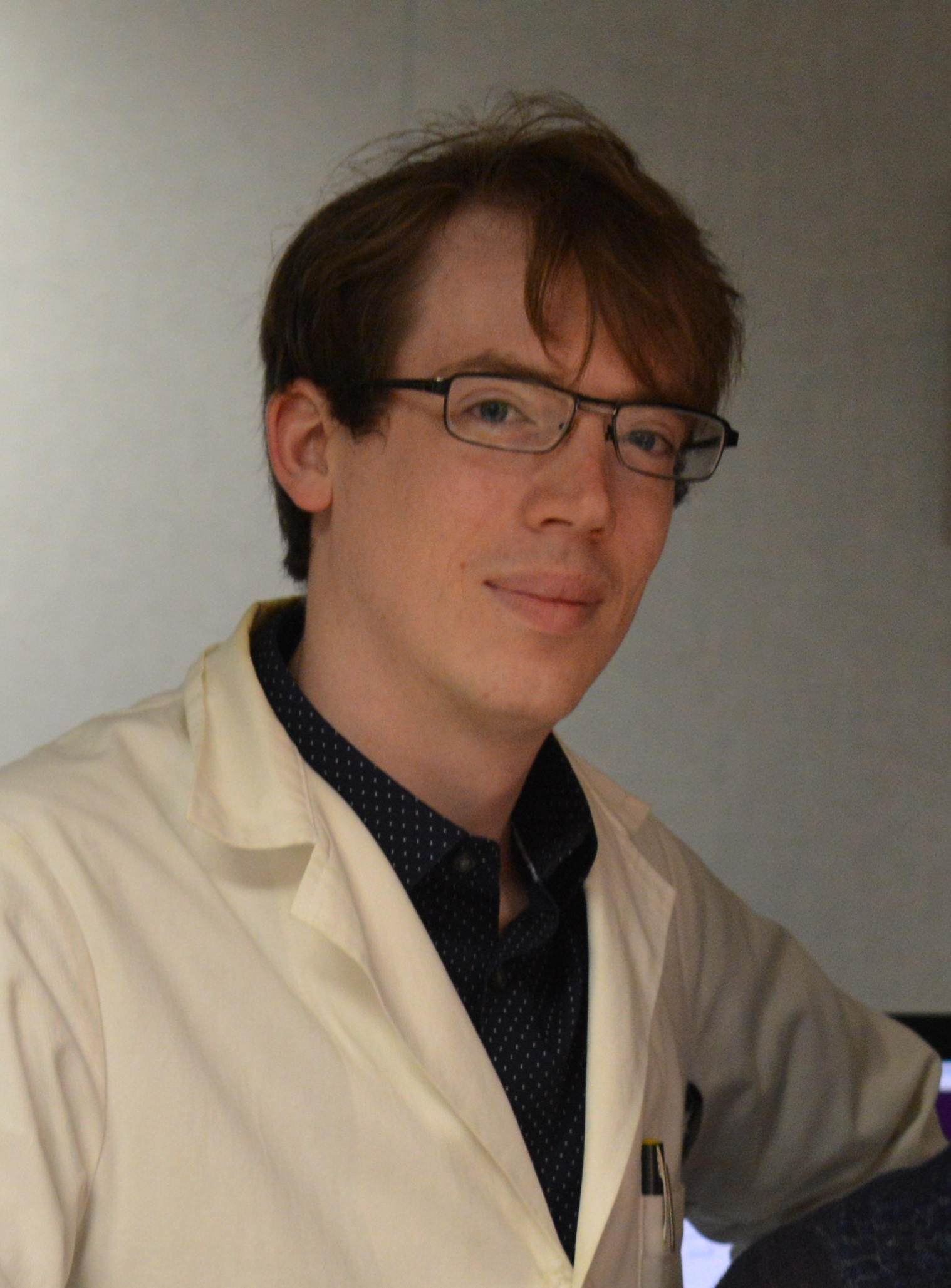

| Jason Kaelber Director, Rutgers New Jersey CryoEM/CryoET Core Facility Associate Research Professor, Institute for Quantitative Biomedicine As director of the cryoEM/ET core facility, I support structural biology research across the IQB and Rutgers community via scientific collaborations, technical services, and the development of new methods. Cryoelectron microscopy can solve structures of biomolecules in their native states at atomic resolution; it is fundamentally a single-molecule imaging technique, and by developing project-specific strategies, we can test hypotheses over a set of individual molecules. Additionally, I investigate the deep evolutionary history of viruses through comparative structural analyses. I have a specific interest in ssDNA viruses including their potential applications.
|
Institute for Quantitative Biomedicine
Room 010D 848-445-5302
|

| Andrew Kern Associate Professor Genetics, School of Arts & Sciences Our research focuses on variation among individuals in the genomes of humans, fruit flies, and plants. In particular we are interested in the role that natural selection has played in shaping patterns of genome variation. Look around you- casual observation of the natural world leads to the intuition that organisms are superbly adapted to their environments. We are interested in dissecting these adaptations at the genomic level, and in turn learning about the nature of the adaptive process. To this end we use computational and mathematical approaches towards using genome sequence data for evolutionary inference.
|
Nelson Biological Laboratories
848-445-3986
|

| Sagar Khare Associate Professor Chemistry & Chemical Biology, School of Arts & Sciences Resident Member, Institute for Quantitative Biomedicine Graduate Program Director, Quantitative Biomedicine The Khare lab seeks to develop new enzymes and proteins using a combination of computational protein design and experimental characterization. Our goal is to develop a quantitative and predictive understanding of enzymatic structures and functions and use this understanding to inform various therapeutic and synthetic applications. Some areas of interest are biodegradation of pollutants, improving cancer chemotherapy, and designing catalytic protein drugs.
|
Proteomics Building Room 208M
848-445-5143
|

| Hossein Khiabanian Assistant Professor Pathology & Laboratory Medicine, Robert Wood Johnson Medical School The research of our laboratory at Rutgers Cancer Institute of New Jersey focuses on cancer genomics and computational biology. We develop novel analytical methods to understand the underlying genetics of human diseases and the molecular epidemiology of disease-causing organisms using high-throughput genomic data. We are interested in identifying prognostic markers in cancer, as well as studying tumor clonal evolution, especially in the contexts of therapeutic resistance, disease transformation, and relapse.
|
Rutgers Cancer Institute of New Jersey
Room 3558 732-235-7445
|

| John Kolassa Professor Statistics & Biostatistics, School of Arts & Sciences Explore approximate and exact statistical methods for small data sets.
|
Hill Center
Room 504 848-445-7674
|
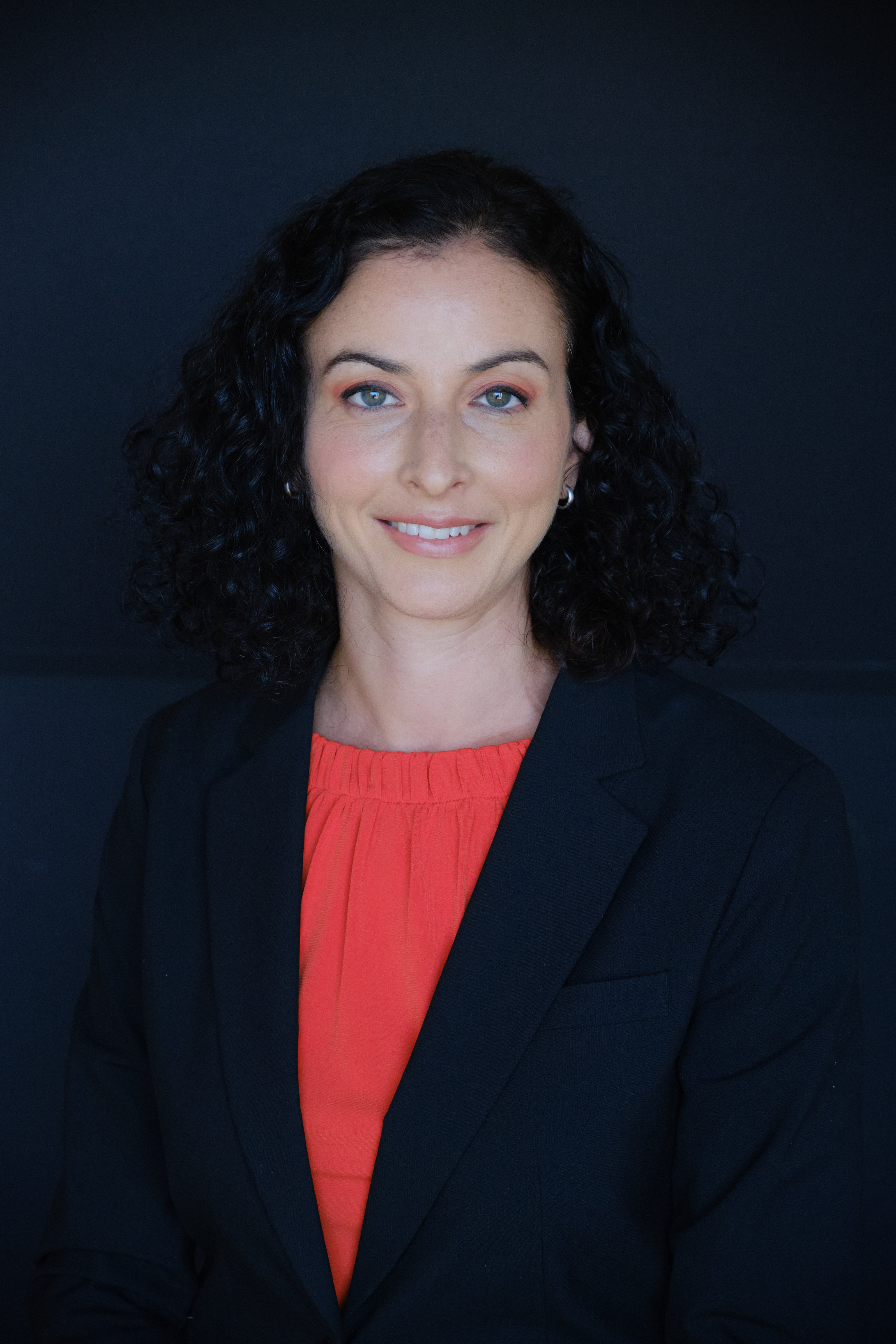

| Anat Kreimer Assistant Professor Biochemistry & Molecular Biology, Robert Wood Johnson Medical School Pediatrics, Robert Wood Johnson Medical School Resident Member, Center for Advanced Biotechnology & Medicine The main focus of our group is to develop predictive models of transcriptional regulation by integrating large-scale datasets to shed light on regulatory processes that are condition-specific. In conjunction with high-throughput experimental techniques, we will leverage such models to identify functional variants and mechanisms of action driving disease. Specifically, we are interested in investigating early stages of neural development using a combination of genomic profiling, functional genomics and computational analysis.
|
Center for Advanced Biotechnology & Medicine 239
347-949-1710
|

| Arek Kulczyk Assistant Professor Biochemistry & Microbiology, School of Environmental & Biological Sciences Resident Member, Institute for Quantitative Biomedicine Our laboratory integrates structural approaches, in particular single-particle cryo-electron microscopy (cryo-EM) and single-molecule techniques to study DNA replication and repair in human mitochondria. We also develop novel correlative light and electron microscopy (CLEM) methods that allow simultaneous visualization of an enzymatic activity and structure determination.
|
Institute for Quantitative Biomedicine
Room 208H 848-445-4553
|

| Edmund Lattime Professor Surgery, Robert Wood Johnson Medical School Biochemistry & Molecular Biology, Robert Wood Johnson Medical School Resident Member, Rutgers Cancer Institute of New Jersey The primary focus of our laboratory is the study of the tumor host interaction with the ultimate gial being the design of effective immunotherapy regimens for cancer. The interaction between the host immune system and tumor is a multifaceted one with the generation of productive immunity requiring the cooperation of immune cells from multiple lineages. Further complicating the system. tumor cells themselves can produce immune suppressive factors.
|
Rutgers Cancer Institute of New Jersey
Room 2010 732-235-8588
|

| Catherine Lawson Research Professor Institute for Quantitative Biomedicine My research explores the diverse landscape of biological structure-function relationships, with the goal of improving our fundamental understanding of life processes. My current projects involve structure database development (EMDataResource, Nucleic Acid Database, Protein Data Bank), with emphasis on improving representation of large biological assemblies.
|
Proteomics Building
Room 109 848-445-5494
|

| Joel Lebowitz George William Hill Professor Mathematics, School of Arts & Sciences Director, Center for Mathematical Sciences Research Mathematical physics and statistical mechanics
|
Hill Center
Room 612 848-445-3117
|

| Jeehiun Katherine Lee Professor Chemistry & Chemical Biology, School of Arts & Sciences Our group combines experimental and computational methods to understand mechanisms of reactions important for chemistry and biology. Specifically, we utilize traditionally physical methods, primarily mass spectrometry and computational chemistry, to tackle problems at the chemistry/biology interface, focusing on catalysis. We work on both biological catalysis, uncovering the mechanisms by which enzymes excise damaged DNA from the genome; as well as organic catalysis, examining N-heterocyclic carbenes as organocatalysts.
|
Wright Rieman Laboratories
Rooms 382 & 372 848-445-6562
|

| Kibum Lee Professor Chemistry & Chemical Biology, School of Arts & Sciences The primary research interest of my group is to develop and integrate nanotechnologies and chemical functional genomics to modulate signaling pathways in stem cells towards specific cell lineages or behaviors. In particular, we are exploring critical problems in cancer research and stem cell biology pertaining to the cell-microenvironmental interactions, and how to control these interactions at the subcellular and single cell level using chemical biology and nanotechnology. From this research effort, we have developed innovative technology platforms that may overcome the critical barriers to harnessing the full therapeutic potential of stem cells for regenerative medicine and personized medicine.
|
Wright Rieman Laboratories
Room 315 848-445-2081
|

| Sang-Hyuk Lee Assistant Professor Physics & Astronomy, School of Arts & Sciences Resident Member, Institute for Quantitative Biomedicine Lee lab is a single-molecule biophysics/biophotonics laboratory. Our lab features a wetlab, cell culture facility, and a one-of-a-kind single-molecule optical instrument. Utilizing advanced fluorescent microscopy, optical tweezers, and digital holography, we are pushing the limit of our understanding of biological processes at the single-molecule level.
|
Proteomics Building Room 308A
848-445-5286
|

| Peter Lobel Professor Biochemistry & Molecular Biology, Robert Wood Johnson Medical School Resident Member, Center for Advanced Biotechnology & Medicine Our laboratory studies lysosomes and associated human. Lysosomes are organelles found in all eukaryotic cells that contain many different proteases, glycosidases, lipases and other hydrolytic enzymes. The lysosome functions as the cell's recycling compartment, breaking down complex biological macromolecules into simple components. The importance of these organelles is underscored by the existence of over thirty human genetic disorders (e.g.. Tay-Sachs disease) where loss of function of a single lysosomal enzyme can lead to severe health problems including neurodegeneration, progressive mental retardation, and early death.
|
Center for Advanced Biotechnology & Medicine
Room 204 848-445-9831
|
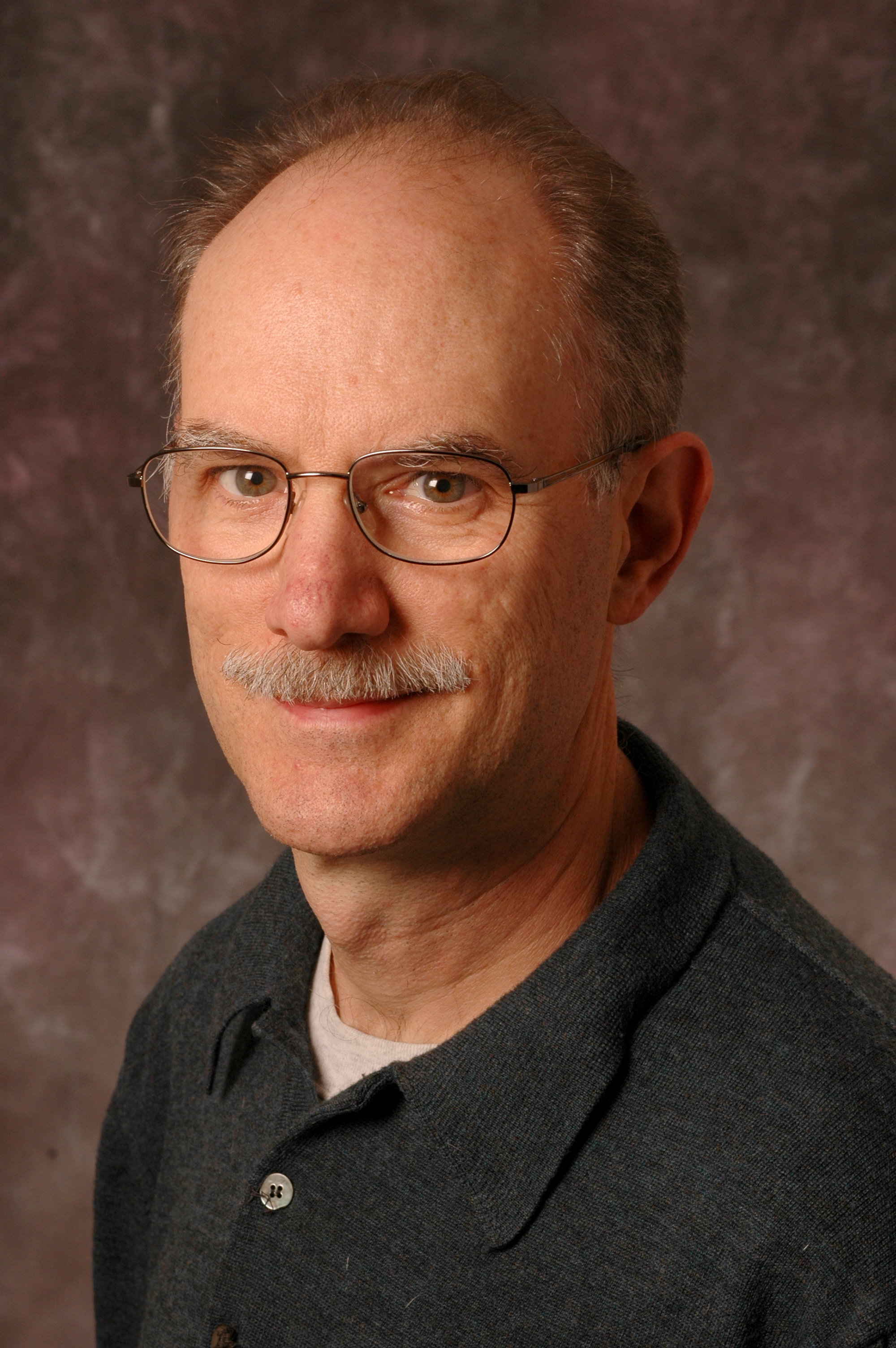

| Richard Ludescher Professor Food Science, School of Environmental & Biological Sciences The research in my lab is concerned with the physical chemistry of edible molecules and biomaterials, with the physical properties, molecular mobility, and molecular states of amorphous solids, and with developing novel applications of luminescence spectroscopy, and especially the identification, characterization and use of edible optical luminescent (fluorescent and phosphorescent) probes, to solve basic scientific as well as practical problems related to the structure, dynamics and function of edible biomaterials.
|
Martin Hall
Room 216 848-932-3516
|

| David M. Lukac Associate Professor Microbiology, Biochemistry & Molecular Genetics, Rutgers New Jersey Medical School Director, Graduate Track in Infection, Immunity & Inflammation We have two major projects: 1. using herpesvirus infection models to investigate the specificity of cellular responses to the Notch signal transduction pathway, and application of learned paradigms to non-viral cancers. 2. investigating the intracellular dynamics of histone deacetylase 6 in autophagy, the stress response, and herpesviral infection.
|
International Center for Public Health
Room E-350 C 973-972-4868
|

| Michael Manhart Assistant Professor Department of Biochemistry and Molecular Biology and Department of Medicine I lead the Quantitative Evolutionary Microbiology Lab. We study how microbes grow, interact, and evolve using a combination of laboratory experiments, computational models, and theoretical approaches. In particular, we seek to understand how fundamental evolutionary processes (such as mutation and selection) shape, and are shaped by, ecological and physiological aspects (such as nutrient limitation and intercellular interactions) of microbial communities. Our long-term goal is to develop a predictive theory of microbiology to solve problems in human health.
|
Institute for Quantitative Biomedicine 306
848-445-9835
|

| Michael B. Mathews Professor & Chair Biochemistry & Molecular Biology, Rutgers New Jersey Medical School Member, Rutgers Cancer Institute of New Jersey His research interests include the control of gene expression, HIV and AIDS, and cellular antiviral defenses. Work in his lab is focused on regulatory systems involved in human disease and host defenses, with RNA-protein interactions as a common theme.
|
New Jersey Medical School-University Hospital Cancer Research Center
Room F1220 973-972-3133
|

| Dimitris Metaxas Distinguished Professor Computer Science, School of Arts & Sciences Physics-based techniques for modeling of organ shape and motion; computer graphics and animation; computational vision; medical imaging
|
Computational Biomedicine Imaging & Modeling Center
Room 11 848-445-2914
|

| Keith Mickolajczyk Assistant Professor Department of Biochemistry and Molecular Biology The Mickolajczyk Lab develops single-molecule biophysical tools including optical tweezers, magnetic tweezers, and high-resolution localization microscopy to study how ATPase motor proteins convert chemical energy into mechanical work. We focus on the enzymology of ribosome biogenesis to uncover the molecular mechanisms underlying this core process, particularly as it related to associated diseases including ribosomopathies and cancer.
|
Robert Wood Johnson Medical School
Public Health Building Room 283 732-235-5888
|

| James Millonig Associate Professor Neuroscience & Cell Biology, Robert Wood Johnson Medical School Senior Associate Dean, School of Graduate Studies My lab studies dorsal CNS development by taking the unique approach of combining mouse genetics with neuroanatomy. Our goal is to identify the pathways, which control the generation and differentiation of dorsal CNS neurons.
|
Center for Advanced Biotechnology & Medicine
Room 238 848-445-9852
|

| Aaron Milstein Assistant Professor of Neuroscience and Cell Biology, RWJMS Resident Faculty at the Center for Advanced Biotechnology & Medicine, RBHS |
|

| Konstantin Mischaikow Distinguished Professor Mathematics, School of Arts & Sciences We use algebraic and computational topological methods to investigate the global structures of multiparameter nonlinear dynamical systems and to study high dimensional spatially-temporally complex data sets.
|
Hill Center
Room 265 848-445-2395
|

| Antonina Mitrofanova Assistant Professor Health Informatics, School of Health Professions Director, Biomedical Informatics Research Her laboratory investigates transcriptional as well as epigenetic mechanisms that drive cancer initiation, progression, and metastases. Furthermore, her lab interests expand to elucidating mechanisms of drug response and resistance and identifying novel drug and drug combinations to target cancer malignancies.
|
Health Informatics
Room 923B 973-972-5241
|

| Prabhas V. Moghe Executive Vice President for Academic Affairs Distinguished Professor Biomedical Engineering The Moghe laboratory has four broad research areas. Two of the areas focus on the design, synthesis, characterization, and mechanisms of interactions of nanoscale materials with cellular matrix proteins. A third area is focused on cell reporter-based biological profiling biological profiling of synthetic polymeric biomaterials for regenerative medicine using high content imaging and high throughput evaluation methods. The fourth effort is on studies of human embryonic stem cell pluripotency and differentiation behaviors on biomaterials.
|
Biomedical Engineering
Room 315 848-445-6591
|

| Gaetano Montelione Professor of Chemistry and Chemical Biology
Constellation Endowed Chair in Structural Bioinformatics Rensselaer Polytechnic Institute The unique strengths of our laboratory involve development of NMR methods and software for analysis of protein structures and dynamics, and in applying these methods in structure-function studies.
|
Jonsson-Rowland Science Center
Rensselaer Polytechnic Institute 110 Eighth Street, Troy, NY 12180-3590 (518) 276-6305
|

| Alexandre Morozov Associate Professor Physics & Astronomy, School of Arts & Sciences Research in my group focuses on understanding how chromatin structure affects gene regulation in eukaryotic organisms. We employ methods from statistical mechanics and polymer physics to develop biophysical models of chromatin. Another major focus area in the group is on mechanisms and theoretical descriptions of molecular evolution, especially on evolutionary constraints due to molecular energetics. Finally, we are interested in non-equlibrium dynamics and propagation on multi-dimensional landscapes and networks.
|
Hill Center
Room 279 848-445-1387
|

| Vikas Nanda Associate Professor Biochemistry & Molecular Biology, Robert Wood Johnson Medical School We design proteins for applications in biomedical research, nanotechnology and use them as tools for understanding protein folding and evolution. Significant progress has been made using computer methods to engineer proteins with novel folds and functions. We maintain and develop software for protein design, structure prediction and docking of protein-ligand complexes.
|
Center for Advanced Biotechnology & Medicine
Room 139 848-445-9810
|

| Joseph Naus Professor Statistics & Biostatistics, School of Arts & Sciences Applied probability, sampling theory, data quality control, clustering and coincidence models, matching in DNA sequences
|
Hill Center
Room 571 848-445-7686
|
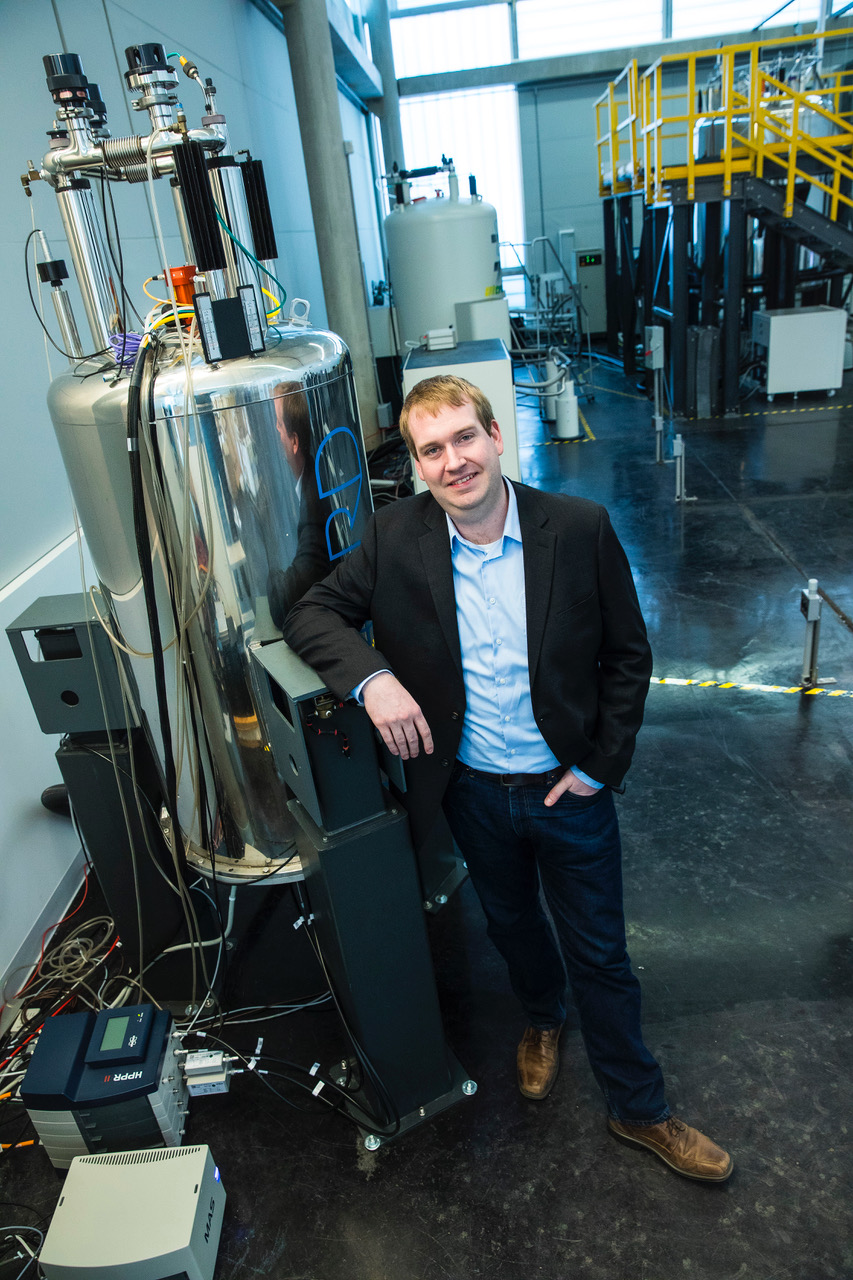

| Andrew Nieuwkoop Assistant Professor Chemistry & Chemical Biology Our research focuses on lipid membranes and their interactions with integral and peripheral membrane proteins. We seek to develop high resolution solid-state NMR (ssNMR) techniques to investigate the structure and dynamics of both proteins and lipid molecules, as well as their interactions. We unitize very-fast magic-angle-spinning to increase the sensitivity and resolution of our spectra, enabling us to address large systems in biologically relevant settings.
|
Wright-Rieman Labs Room 311
848-445-2626
|

| Wilma Olson Mary I. Bunting Professor Chemistry & Chemical Biology, School of Arts & Sciences All of the processes necessary for the survival of a living system hinge on its ability to store and read the genetic information encoded in its DNA. The packaging of the long human genome into the very small confines of the nucleus is complicated by the necessity of maintaining the accessibility of the DNA for genetic processing. The Olson group is establishing new chemistry- and physics-based computational methodologies to unravel the mystery of how genomes can at once be tightly packed and yet available for read-out.
|
Wright Rieman Laboratories
Room A209B 848-445-3993
|

| Zhiping Pang Assistant Professor Neuroscience & Cell Biology, Robert Wood Johnson Medical School My laboratory studies the neural basis of the regulation of feeding, satiety, metabolism and obesity. Our studies may provide insights into the neural causes and consequences of childhood obesity. We also developed novel techniques for deriving neuronal cells from primary skin cells and pluripotent stem cells, providing novel opportunities to study the pathogenesis of neurological disorders, including pediatric developmental disorders and autism spectrum disorders.
|
Child Health Institute of New Jersey
Room 3277 732-235-8074
|

| Manish Parashar Visiting Professor Computer Science, School of Arts & Sciences Founding Director, Rutgers Discovery Informatics Institute Director, The Applied Software Systems Laboratory High-performance applied, distributed, and autonomic computing; computational and data-intensive science and engineering
|
Hill Center
Room 628 848-455-5388
|

| Smita Patel Professor Biochemistry & Molecular Biology, Robert Wood Johnson Medical School The overarching goal of our research is to understand the basic mechanisms of life's essential processes including DNA replication, DNA transcription, and pathogen recognition by innate immune receptors.
We specialize in quantitative characterization of protein-nucleic acids interactions and dissection of multistep enzymatic reactions into its elementary steps for complete characterization of biochemical pathways. We are pushing fluorescence based methods, transient state kinetics, single molecule kinetics, and computational kinetic modeling to characterize the mechanisms of nucleic acid motor proteins such as DNA and RNA helicases and polymerases in various processes.
Our current research investigates the cooperativity and enzymatic mechanisms of multisubunit protein complexes that catalyze human mitochondrial DNA replication and transcription and self and non self RNA recognition by the RIG-I like helicase innate immune receptors. Our research has application in biotechnology and therapeutics.
|
Robert Wood Johnson Medical School Research Building/School of Public Health
Room 283 732-235-3371
|

| Vladimir Pavlovic Professor Computer Science, School of Arts & Sciences Sequence Analysis and Modeling Lab conducts research in computational modeling methods for analysis of multimodal and multivariate data, with particular focus on sequential data, such as time-series (video, audio, motion sensors), biomedical data (imaging, genomic, proteomic, etc.) We develop mathematical and statistical models and the accompanying algorithms that can rapidly process large and heterogeneous datasets and reveal the underlying, latent factors affecting the data.
|
Computational Biomedicine Imagining & Modeling Center
Room 006 848-445-8846
|

| Ezra Peisach Assistant Research Professor Chemistry & Chemical Biology, School of Arts & Sciences Biocurator, RCSB Protein Data Bank My research centers on the automation of biocuration activities, distributed computing and distributed infrastructure management of distributed of RCSB Protein Data Bank.
|
Proteomics Building
Room 118 848-445-4948
|

| John Pintar Professor Neuroscience & Cell Biology, Robert Wood Johnson Medical School Graduate Program Director The Pintar laboratory has used mammalian molecular genetic approaches to produce 16 lines of mice containing null alleles for genes involved in growth as well as nervous system development and function. Current work involves both the study of these mutant strains using multiple analytic levels as well as producing conditional knock-outs for several genes. Perhaps most notably, the lab has produced knock-outs of the entire opioid system and is now studying the consequences of these mutations on pain perception and anxiety as well as on drug-induced tolerance and dependence.
|
Robert Wood Johnson Medical School Research Building/School of Public Health
Room 359 732-235-4250
|

| Vincenzo Pirrotta Professor and Chair Molecular Biology & Biochemistry, School of Arts & Sciences Associate Member, Rutgers Cancer Institute of New Jersey Chromatin structure and dynamics. polycomb silencing mechanisms, epigenetic mechanisms, developmental gene regulation, genomic programming, Drosophila genetics
|
Nelson Biological Laboratories
Room A121 848-445-2446
|

| Arnold Rabson Laura Gallagher Endowed Chair, Developmental Biology Pediatrics; Pharmacology; & Pathology & Laboratory Medicine, Robert Wood Johnson Medical School Director, Child Health Institute of New Jersey Full Member, Cancer Institute of New Jersey The Rabson laboratory studies the pathogenesis of human retroviral infections including the human T cell leukemia virus type 1-an important cause of human leukemias and inflammatory diseases, and the human immunodeficiency virus. He also studies the roles of the NF-kB transcription regulators in the genetic basis of normal and malignant T cells, and the molecular basis of hematopoiesis and childhood leukemias.
|
Child Health Institute of New Jersey
Room 3210 732-235-9523
|

| Fred Roberts Distinguished Professor Mathematics, School of Arts & Sciences Director Emeritus, DIMACS Director, Homeland Security Center of Excellence CCICADA Mathematical modeling of problems in the biological, social, and environmental sciences; biological and medical data; social science aspects of biomedicine and epidemiology; metrics and measurement; applications of discrete mathematics; information-based decision making
|
Center for Discrete Mathematics & Theoretical Computer Science
848-445-4303
|

| Charles Roth Professor Chemical and Biochemical Engineering, School of Engineering Biomedical Engineering, School of Engineering The Roth laboratory applies approaches from nanobiotechnology to biomedical problems, ranging from cancer to heterotopic ossification. Our main expertise is in the development of multifunctional nanoparticles to overcome delivery barriers ranging from the systemic to intracellular, and their application for gene silencing (antisense, RNAi), drug delivery and biomedical imaging.
|
Chemical & Biochemical Engineering;
Biomedical Engineering Room 205 848-445-6686
|

| Monica Roth Professor Pharmacology, Robert Wood Johnson Medical School Full Member, Rutgers Cancer Institute of New Jersey Our research studies three stages of the retroviral life-cycle: entry, replication and integration. Retroviruses are RNA viruses capable of inducing various diseases including cancers and immunodeficiencies. The focus of our research is to study three stages in the infectious retroviral life cycle. These include the entry, replication and integration of the virus.
|
Robert Wood Johnson Medical Research Tower
Room 636 732-235-5048
|

| Anirvan Sengupta Professor Physics & Astronomy, School of Arts & Sciences I have two main broad subjects of interest: Condensed Matter Physics and Cellular Biology.
In condensed matter physics, I have been working on effects of interaction and disorder on electrons. I am specially interested in magnetic phase transitions in metals. More technically speaking, I have tried to use insights gained from impurity models like Kondo model to understand the local physics of extended interacting systems.
I have been interested in off-equilibrium dynamics of unidirectional processes, specially in context of ion pumping on cellular membranes. I am also interested in enzymatic networks and how they control gene expression or transduction.
|
Serin Physics
Room E263 732-445-4668
|

| Konstantin Severinov Professor Molecular Biology & Biochemistry, School of Arts & Sciences Principal Investigator, Waksman Institute of Microbiology Transcription is the central step, and a major regulatory checkpoint of gene expression. Defective transcription regulation is the common cause of aberrant growth and development and may result in malignant transformation. Transcription is carried out by DNA-dependent RNA polymerases–large, multisubunit molecular machines. Understanding RNA polymerase (RNAP) structure and function is a key to understanding gene expression in molecular detail. The long-term objective of our research is to uncover the molecular basis of transcription mechanism and regulation through structure-functional analysis of bacterial RNAP and associated proteins. In addition, we use bacteriophage development as a model system to study temporal regulation of gene expression and to uncover novel mechanisms of transcription regulation. We also study microcins, small ribosomally-synthesized inhibitors of bacterial growth.
|
Waksman Institute of Microbiology
Room 224 848-445-6095
|
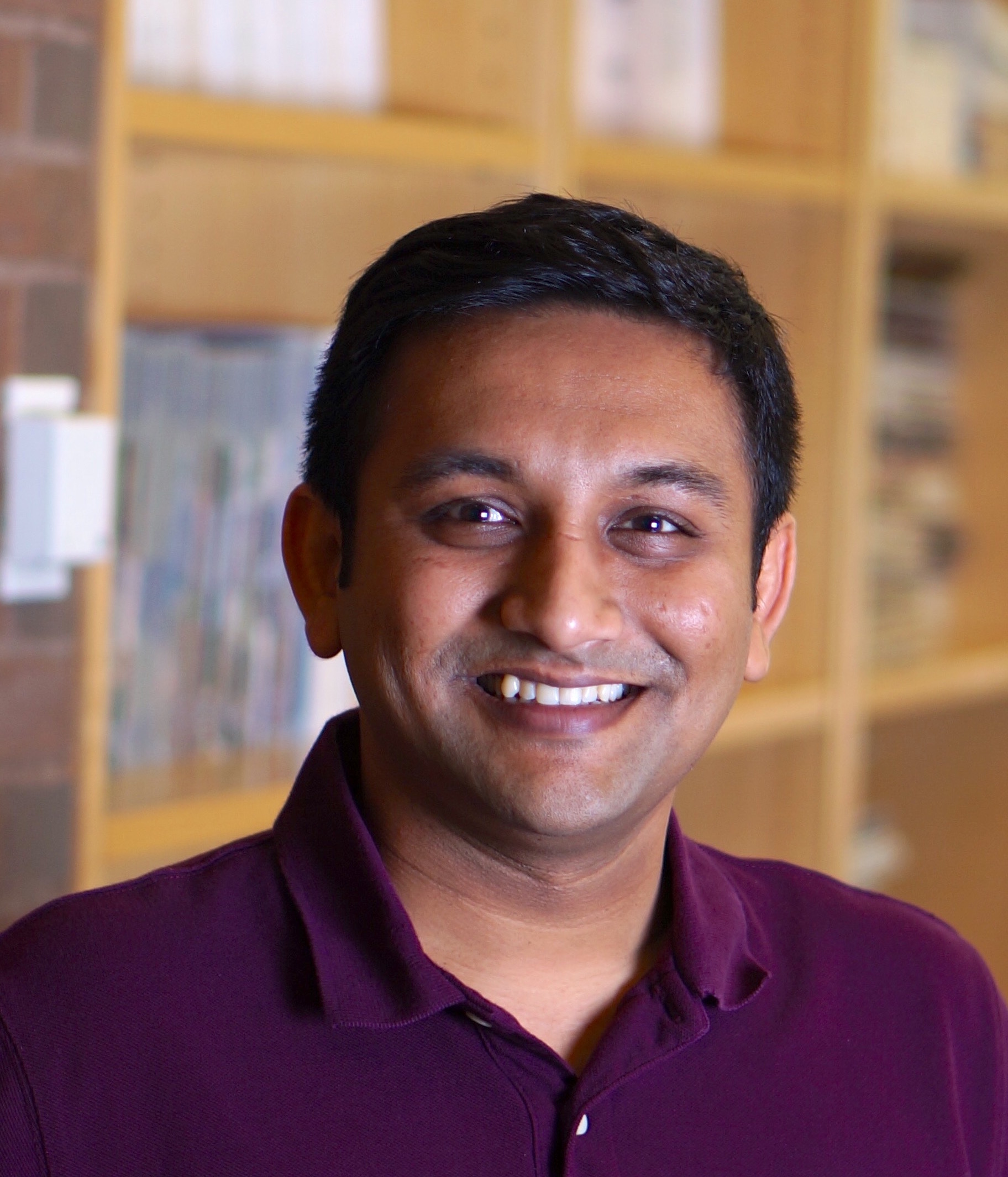

| Premal Shah Assistant Professor Genetics, School of Arts & Sciences Our research integrates mechanistic models of biological processes with population-genetic models to understand the forces shaping genome architecture. Our research program is focused on: evolution of codon usage bias and protein translation (by building detailed biophysical models of protein translation, we seek to explain the mechanistic bases of protein production in a cell as well as the factors that drive the evolution of biased codon usage) and epistasis in protein evolution (by combining models of protein structure and function with a population-genetic framework, we seek to explain how mutations individually and collectively influence protein evolution)
|
Life Sciences Building
Room 326 848-445-9664
|
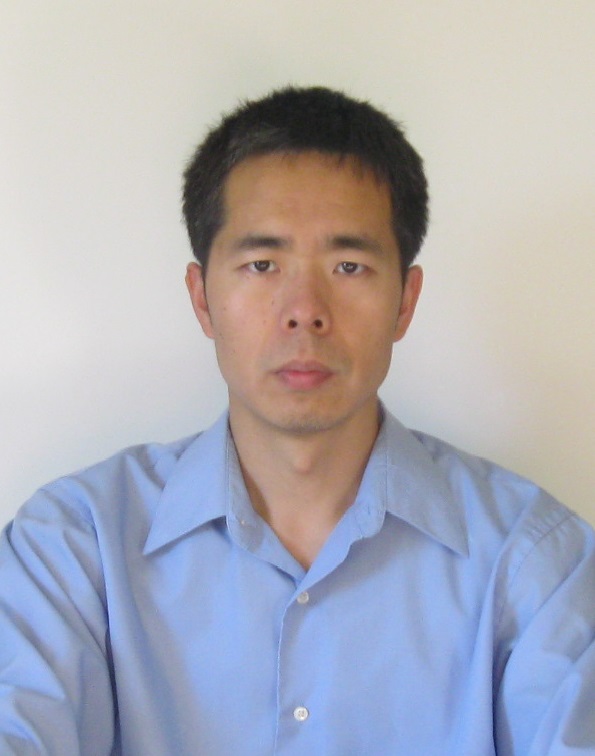

| Chenghua Shao Assistant Research Professor Chemistry & Chemical Biology, School of Arts & Sciences My research focuses on data curation and analysis at the Protein Data Bank (PDB). PDB provides free public deposition and distribution services on macromolecular 3D structures determined by experimental methods. As PDB data are broadly used in biomedical research, we develop system and methods for data process and validation, to ensure data accuracy and archive consistency. With combination usage of structure biology, biochemistry, and statistics, we also perform data analysis to find trends, patterns, outliers, and the underlying biological explanation, which enables better data collection and curation to improve data quality. In addition, I supervise curation on the structural and chemical data of ligands and drugs to ensure the PDB representation of such compounds is up to the standard for their usage in biomedical research.
|
Proteomics Building
Room 118 848-445-4968
|

| Troy Shinbrot Professor Biomedical Engineering, School of Engineering Our first area of research concerns the formation of structures during development of tissues and organs, especially in the central nervous system. This work includes the study of neuronal pathfinding following spinal cord injury and a simulation of the development of folds, for example in the mammalian cerebellum. A second area of investigation deals with the flow and mixing of both dry and wet grains. The results have applications spanning a broad range from pharmaceutic formulation to geophysical pattern formation.
|
Biomedical Engineering
Room 310 848-445-6584
|

| David Shreiber Professor & Chair Biomedical Engineering, School of Engineering We are interested in understanding Central Nervous System injury, repair, and regeneration. We study and attempt to dictate the tissue and cellular biomechanics associated with traumatic injury using in vivo, in situ, and computational models. We also use in vitro and computational models to investigate mechanotransduction as a mechanism underlying the response to acupuncture.
|
Biomedical Engineering
Room 312 848-445-6589
|

| Eduardo Sontag Distinguished Professor Department of Bioengineering Department of Electrical and Computer Engineering Northeastern University Our interests cover a wide variety of areas, including systems and synthetic biology, modeling in cancer and immunology, feedback control and dynamical systems, computational complexity and theory of computing, & drug combinations and resistance
|
Northeastern University
805 Columbus Ave, Boston, MA 02118 848-445-4552
|

| Ruth Steward Professor Molecular Biology & Biochemistry, School of Arts & Sciences Principal Investigator, Waksman Institute of Microbiology We are studying the mechanisms controlling RNA processing (i.e. translation, localization, and stability)of specific sets of transcripts. We have identified and characterized a novel highly conserved gene, Zfrp8/PDCD2 and shown that it is essential in stem cells in flies, mouse and human and is also required for growth of cancer cells. Recently, we discovered that it is required for the nuclear export of select mRNAs and TE transcripts. It also interacts with the small ribosomal subunit and complexes with mRNA binding proteins. Our data suggest that Zfrp8/PDCD2 controls subcellular localization of select RNAs and the association of mRNA-RNP with ribosomes.
|
Waksman Institute of Microbiology
Room 252 848-445-3917
|
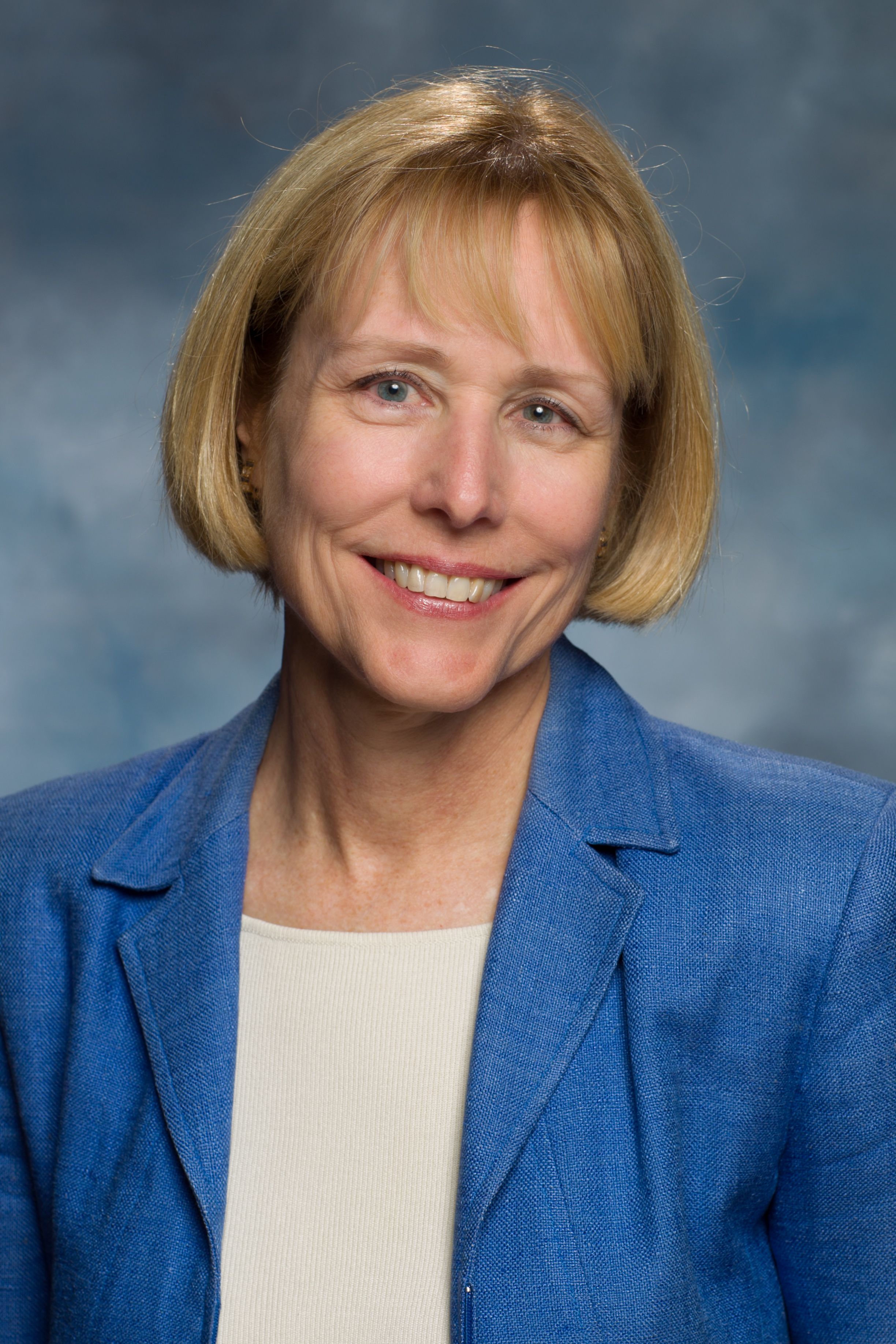

| Ann Stock Professor Biochemistry & Molecular Biology, Robert Wood Johnson Medical School Interim Director, Center for Advanced Biotechnology & Medicine Research in the Stock laboratory focuses on investigating the molecular mechanisms of receptor-mediated signal transduction. Studies range from elucidating structure/function relationships in signaling proteins in vitro using a combination of molecular genetic, biochemical, and X-ray crystallographic methods to systems biology approaches aimed at characterizing signaling pathways in vivo. Specific interest is directed toward investigating the role and regulation of covalent modifications of proteins in bacterial signaling pathways.
|
Center for Advanced Biotechnology & Medicine
Room 106A 848-445-9812
|
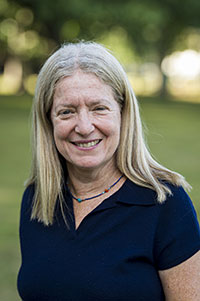
| Judith Storch Distinguished Professor Nutritional Sciences, School of Environmental & Biological Sciences Campus Dean, G.H. Cook Campus
Co-Graduate Program Director Lipids such as fatty acids and cholesterol are involved in innumerable cellular processes, including energy storage and production, membrane biogenesis, signal transduction, and the regulation of gene expression. Nevertheless, the mechanisms by which lipids are transported and targeted within cells remain largely unknown. Abnormal lipid trafficking, such as that occurring in lipid-storage diseases, can lead to severe cellular pathologies. The overall focus of research in this laboratory is on lipid traffic in cells, with particular emphasis on long-chain fatty acids, monoacylglycerols, and cholesterol.
|
Food Science & Nutritional Sciences Building
Room 311-D 848-932-1689
|

| Zoltan Szekely Research Professor Pharmaceutics, Ernest Mario School of Pharmacy Director, Chemical Biology Core Facility Associate Member, Rutgers Cancer Institute of New Jersey Antibody-drug conjugates, focusing on linkers and drugs, Tumor hypoxia and its application for prodrug design, Sequence selective DNA-targeting agents, Drug delivery by nanoparticles and bio-conjugates, Drug design, High-throughput synthesis of small molecules and peptides
|
William Levine Hall
Room 225 732-325-5851
|

| Bin Tian Professor and Program Co-Leader, Gene Expression & Regulation Program, Ellen and Ronald Caplan Cancer Center Co-director, Center for Systems & Computational Biology Wistar Institute Our research is focused on understanding how gene expression is regulated at the RNA level. My lab was among the first to discover the widespread nature of alternative polyadenylation (APA) using bioinformatic and genomic approaches.
Wistar Institute
|
New Jersey Medical School-University Hospital Cancer Research Center
Room F1230 973-972-3615
|

| Jay Tischfield Duncan & Nancy MacMillan Distinguished Professor Genetics, School of Arts & Sciences Pediatrics & Psychiatry, Robert Wood Johnson Medical school Founder, CEO and Scientific Director, RUCDR Infinite Biologics Executive Director, Human Genetics Institute of New Jersey Jay Tischfield's group is investigating the genetics and pathophysiology of complex neuropsychiatric disorders such as Tourette Syndrome and addiction disorders such as alcoholism. Other research areas include genomic structure and modifications associated with cell differentiation and maintenance of genome stability with emphasis on mechanisms of loss of heterozygosity.
|
Life Sciences Building
Room 136 848-445-1027
|

| Brinda Vallat Associate Research Professor Institute for Quantitative Biomedicine PDB-Dev Project Lead I am a computational biologist interested in understanding the molecular basis of life. My research interests include modeling and analysis of biomolecular structure and function using data obtained from experimental and theoretical methods. I currently lead the PDB-Dev project, which is focused on building the infrastructure for archiving integrative structural models of macromolecules. Integrative modeling methods combine complementary information from various experimental and computational techniques to determine the structures of complex macromolecular assemblies. The complexities of handling different data types involved in integrative modeling provide challenging opportunities for data acquisition, standardization, archiving, validation and dissemination.
|
Proteomics Building
Room 105 848-445-8377
|

| Andrew Vershon Professor Molecular Biology & Biochemistry, School of Arts & Sciences Principal Investigator, Waksman Institute of Microbiology Director, Waksman Student Scholars Program A major focus of his research is on the regulation of transcription in the yeast, Saccharomyces cerevisiae. Specifically, he is investigating how different regulatory proteins interact to control gene expression and how these interactions influence the regulatory activity of the proteins. As Professor, Vershon is one of the most enthusiastic molecular biologists, and loves involving undergraduates in research . During the summer, Dr. Vershon works with high school teachers to encourage their students to participate in genuine molecular biology research in the Waksman Summer Scholars Institute program.
|
Waksman Institute of Microbiology
Room 234 848-445-2905
|

| Michael Verzi Assistant Professor Genetics, School of Arts & Sciences Member, Rutgers Cancer Institute of New Jersey Member, Human Genetics Institute We employ epigenomic approaches to understand how the mammalian intestine develops and functions, and to understand why this tissue is susceptible to cancer and inflammatory disease. We put particular emphasis on transcriptional regulatory mechanisms governing these processes.
|
Life Sciences Building
Room 127 848-445-9578
|

| Lu Wang Associate Professor Chemistry & Chemical Biology, School of Arts & Sciences Resident Member, Institute for Quantitative Biomedicine We develop and utilize theoretical and simulation methods to study the structure, dynamics and spectroscopy of biological systems.
|
Proteomics Building Room 208D
848-445-4555
|

| Sijian Wang Professor Statistics & Biostatistics, School of Arts & Sciences Resident Member, Institute for Quantitative Biomedicine Our long-term research goal is to develop, evaluate, and disseminate effective statistical and computational methods to identify the biological mechanisms underlying complex diseases, and use the information to improve patient outcomes. We utilize advanced statistical and machine learning methods (including high-dimensional regression, causal inference, modern survival analysis, SVMs, boosting, random forest, deep learning and reinforcement learning) to addresses some of the most important statistical and computational deficiencies that are currently preventing the scientific community from turning valuable high-throughput measurements into meaningful results.
|
Institute for Quantitative Biomedicine
Room 308D 848-445-4562
|

| William Welsh Norman H. Edelman Professor Department of Pharmacology, Robert Wood Johnson Medical School Associate Director, Division of Cheminformatics, Biomedical Information Shared Resource, RCINJ Dr. Welsh's research program specializes in the development and application of computational algorithms and tools to accelerate and streamline drug discovery.
|
Research Tower
Room 561 609-577-4413
|

| Jinchuan Xing Assistant Professor Genetics, School of Arts & Sciences My long-term research interest is to understand the mechanisms and consequences of human genomic variation. My current projects involve elucidating human population history and genetic adaptation at both global and regional scale, with or without disease implication.
|
Life Sciences Building
Room 325 & 330A 848-445-9663
|

| Martin Yarmush Paul & Mary Monroe Chair Biomedical Engineering, School of Engineering The research activities in our laboratory address various challenging areas in biotechnology and bioengineering. Among the current projects are: the development of new nanoparticle technology to enhance wound healing and siRNA delivery; microfabricated tissue-on-a-chip-systems for drug and environmental toxin testing; pulsed electric field techniques to promote scarless wound healing and wound disinfection; whole liver re-engineering through recellularization of decellularized scaffolds and revitalization perfusion of marginal organs; supercooling preservation of cells, tissues, and organs; encapsulated mesenchymal stem cells for treatment of spinal cord injury and osteoarthritis; and development of automated robotic venipuncture devices with point-of-care capabilities. Success in tackling these projects is enabled by the use of state-of-the-art techniques that include microfabrication and nanotechnology; physical biochemistry; genomics, proteomics and genetic engineering; cell biology and tissue engineering; advanced microscopic imaging; physiologic instrumentation; animal studies; and numerical simulation.
|
Biomedical Engineering
Room 231 848-445-6528
|

| Darrin York Professor Chemistry & Chemical Biology, School of Arts & Sciences Resident Member, Institute for Quantitative Biomedicine The York Group research involves the development of a broad range of theoretical methods aimed, ultimately, at providing the computational biology community with greatly improved tools for the simulation of biological macromolecules in solution. The main application focus of our group is on an extremely interesting and challenging area of biology: the study of the molecular mechanisms of RNA catalysis. This area is immensely important, not only from a fundamental biological perspective, but also for the design of medical therapies that target genetic disorders, and new biotechnology such as allosteric RNA chips.
|
Proteomics Building Room 308G
848-445-5199
|

| Jasmine Young Research Professor Chemistry & Chemical Biology, School of Arts & Sciences Biocuration Team Lead, RCSB Protein Data Bank Global Project Lead, worldwide Protein Data Bank I oversee practices, management, and OneDep tool developments for data deposition, curation, validation, and remediation of the Protein Databank (PDB) archive to ensure data standardization, high quality and uniformity. Collaborating with wwPDB partners globally, I represent RCSB PDB to set common annotation practices and procedures, set data standards, and establish daily operation and weekly release procedures across wwPDB sites. I also manage wwPDB projects across wwPDB partners, including development of OneDep tools, data remediation, data analysis, and strategic planning.
|
Proteomics Building
Room 111 848-445-4920
|

| Miguel Zaratiegui Associate Professor Molecular Biology & Biochemistry, School of Arts & Sciences Full Member, Rutgers Cancer Institute of New Jersey Role of transposons and heterochromatin in genome regulation and stability
|
Nelson Biological Laboratories
Room A133 848-445-1497
|

| Steve Zheng University Professor of Pharmacology Robert Wood Johnson Medical School Co-Director, The Cancer Pharmacology & Pre-clinical Therapeutics Program Resident Member, Rutgers Cancer Institute of New Jersey Growth is the process whereby cells accumulate mass, which determines the sizes of cells, organs and organisms. Deregulation of growth is directly linked to human diseases from cancer to aging. Specifically, our current focus is on the target of rapamycin (TOR) protein, a highly conserved phosphatidylinositol kinase-related kinase (PIKK) and a central controller of cell growth. We are interested in dissecting the basic signal transduction mechanisms by TOR and in understanding the mechanisms by which cancer cells resist rapamycin and rapamycin analogs. We hope to ultimately develop more effective therapeutics against human cancer.
|
Rutgers Cancer Institute of New Jersey
Room 3549 732-235-6879
|
